Computed Structures of Core Eukaryotic Protein Complexes
January 25, 2022 | Terry Sharrer
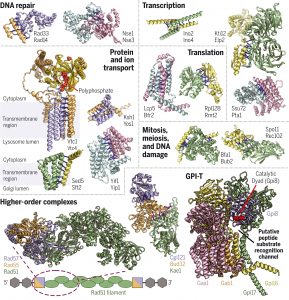
Computed structures of core eukaryotic protein complexes
“We [a wide collaboration of thirty researchers] take advantage of advances in proteome-wide amino acid coevolution analysis and deep-learning-based structure modeling to systematically identify and build accurate models of core eukaryotic protein complexes within the Saccharomyces cerevisiae proteome. We use a combination of RoseTTAFold and AlphaFold to screen through paired multiple sequence alignments for 8.3 million pairs of yeast proteins, identify 1,505 likely to interact, and build structure models for 106 previously unidentified assemblies and 806 that have not been structurally characterized.” MORE
Image Credit: Science.org